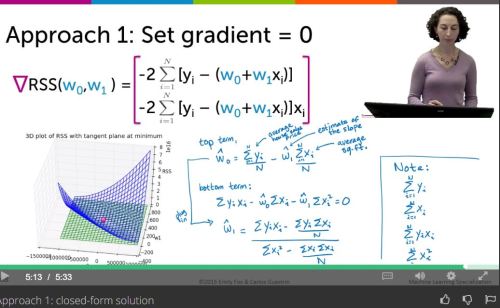
Machine Learning – Stanford -Week 8 – Clustering and PCA
Clustering – K-means algorithm
Continuing with Andrew Ng’s course and finally finished week 8 and delivered homework before deadline. So here again just to note some key points.
K-means
Basically speaking the K-means method is very simple and strait forward, one only need to repeat 1. assigning all samples to one of the closest centroids; 2. move centroids to the middle of those assigned samples. However the implementation is a little tricky to make correct due to the indexing.

And it is worth to mention that not only well separated clusters, but also a group of connected well (or non-separated clusters) can be also clustered via K-means method. This can be particularly useful, for example, a T-shirt company would like to only make S, M and L size of T-shirt for all people with different height and weight.

And here is a Pop-quiz:

More about random initialization


We could imagine that, when k is relatively small (for example k in range of 2 to 10), there is a big chance that the random initialization will put some centroids very close to each other in the beginning and result in some local optima.
The way to solve this issue is also very simple – do clustering (with random initialization) multiple times and then select the centroid group with the lowest cost.

Choosing the value of K
Another question is how should we choose or decide how many clusters should there be for our data. Andrew suggested two method:
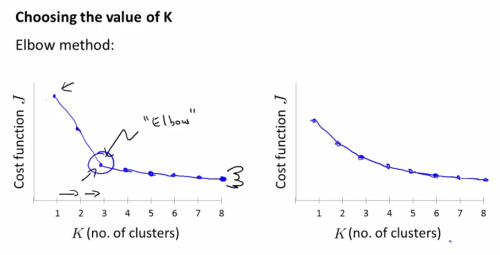
- Elbow method, which is running the K from 1 to many and if we could see a clear “elbow” shape of cost curve, we can choose the K of that elbow point.
- Based on practical or business/application specification.


Okay, so here are some pop quiz for K-means clustering:





And here I would like to add a image compression result made from the homework result – so for this example we used K-means clustering to cluster all pixels of a image with only 8 colors, and the original image is in RGB color of 256 colors. With K=8 we could use the algorithm to cluster all pixels with similar colors into one of the 8 colors. Notably that the value of the 8 colors are not pre-defined, they will gradually become some of the major colors on the picture (while the “moving centroids step” in the k-means algorithm). After the clustering is done, we can simply do the compression by finding the nearest centroid of every pixel and then assign that pixel of that color of centroid. Since we only have 8 colors, a 3 bit storage room for each pixel will be enough, while the 8 centroids are holding a RGB color each as the “cluster center” color 🙂
Principal Component Analysis – PCA
The motivation of principal component analysis is mainly about data compression – for example, how to casting a high dimension data down to a lower dimension space. This will help saving space of data storage, also improving certain algorithm speed, and also to help visualization since a data more than 3D, 4D will become impossible to be visualized directly.


So the so-called Principal Component Analysis (PCA) is just a method of reduce high dimension data to lower dimension data.
One thing we should notice is that even though it looks similar, but PCA and linear regression are two different algorithms and the goal is also totally not the same. For linear regress we have a value y we would like to predict, however for PCA all data are treated the same, there is no special label value.
PCA – How to
As Andrew mentioned, the mathematical proof of the following way of doing PCA is very difficult and also beyond the scope of the course, so here we just need to know how to use it instead. It turned out to just do a dimension reduction is actually very simple. Two lines of matlab code would be enough. The key is to utilize some build in “svd” function.

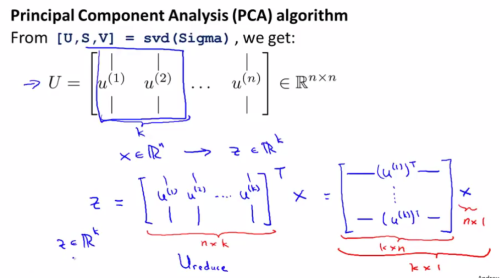
And a summary of the stuff above: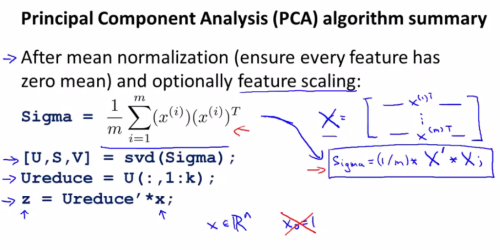
Again, here a pop quiz
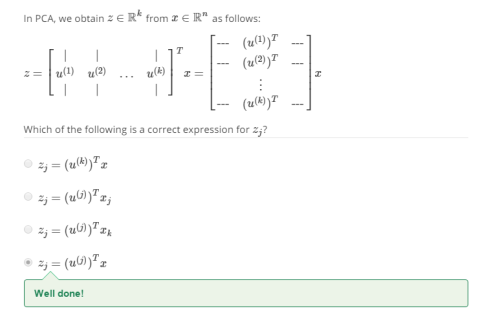

Reconstruction


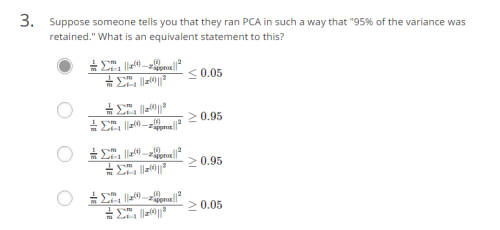
How to choose which level of dimension k we can reduce to?



Another quiz:
PCA – What can I use if for?




And some final pop quiz for under standing the topic better: